Dan Milkie
I am a Senior Scientist in Nobel Laureate Eric Betzig’s group at the Janelia Research Campus. My interests are in tool building for science, including super resolution light microscopy and adaptive optics.
Cells did not evolve to live on the glass coverslip nor under the burning intensity of high power lasers. Unlocking what cells and biology are doing dynamically and in their native environment is key to understanding how complex biologic systems work.
I create the microscope hardware and software control systems for the Betzig Group and others, and bring to bear any new technology we can leverage (FPGA, Spatial Light Modulators, Deformable Mirrors, Piezo stage, >GB/s cameras,… ).
I build, first, for the hands of power users. Next comes a large investment: taking their achieved wisdom and baking it into new hardware and software versions, but it’s totally worth it. The pay-off comes with wide impact distribution, commercialization, and use by non-specialists. Tech can’t be a limited to a unicorn machine in a single lab doing a hero experiment.
Volumetric imaging microscopy platforms deliver a firehose of 4D data, which generate a host of new, enticing challenges (data transfer, storage, curation, processing, visualization, ML analysis) which are the catnip to my future.
Download my resumé.
View my publications.
- Microscopy
- Machine Vision
- LabVIEW
- Python
- Automation
- Efficiency
PhD in Physics, 2008
University of Pennsylvania
BSc in Physics, 2002
The College of William and Mary
Experience
Created the hardware control system and control software for all Betzig microscopes and few others
- Lattice Light Sheet
- AO Lattice Light Sheet
- AO Bessel
- Structured Illumination Micrsocope
- Opposing Objective Microscope
- Cryo-SIM/-PALM
- MOSAIC (Adaptive Optics added to everything we can in one scope)
Grew a small machine vision-centric company, while solving industrial part-inspection tasks and scientific research automation challenges.
- Built systems to inspect: Dinnerware, truck brakes, agriculture seeds, optical fiber, mass flow controllers,…
- Point-cloud 3D laser height mapping, bright & dark field imaging, Area scan cameras, and Conveyor-based line-scan camera systems
- Oversaw projects, contracts, budgets, sales, small teams, recruitment, and timelines…and other the many hats that come with a small business.
Studied and measured carbon nanotubes using optical, electrical, and thermal measurements, with some electron microscopy and magnetic measurements along for the ride too.
Multi-day measurement averaging and mapping of large parameter space required heavy (and robust!) automation. I learned and loved to build control algorithms for collecting and analyzing these data automatically, and got a big kick out of bringing that over to a multi-user facility.
Featured Publications
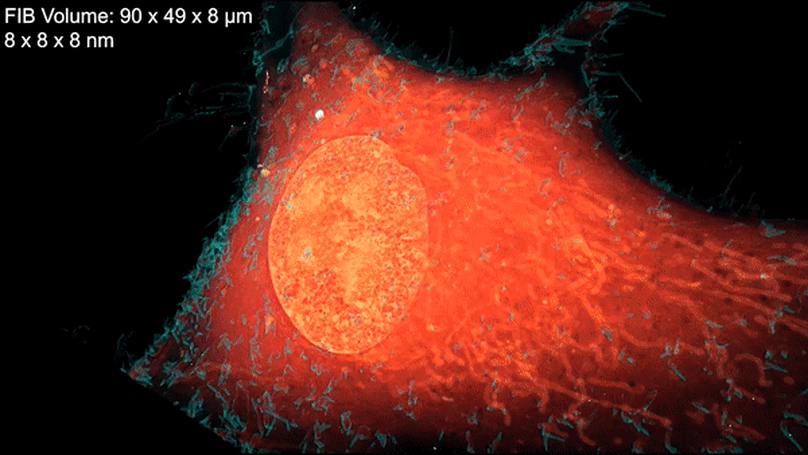
Within cells, the spatial compartmentalization of thousands of distinct proteins serves a multitude of diverse biochemical needs. Correlative super-resolution (SR) fluorescence and electron microscopy (EM) can elucidate protein spatial relationships to global ultrastructure, but has suffered from tradeoffs of structure preservation, fluorescence retention, resolution, and field of view. We developed a platform for three-dimensional cryogenic SR and focused ion beam-milled block-face EM across entire vitreously frozen cells. The approach preserves ultrastructure while enabling independent SR and EM workflow optimization. We discovered unexpected protein-ultrastructure relationships in mammalian cells including intranuclear vesicles containing endoplasmic reticulum-associated proteins, web-like adhesions between cultured neurons, and chromatin domains subclassified on the basis of transcriptional activity. Our findings illustrate the value of a comprehensive multimodal view of ultrastructural variability across whole cells.
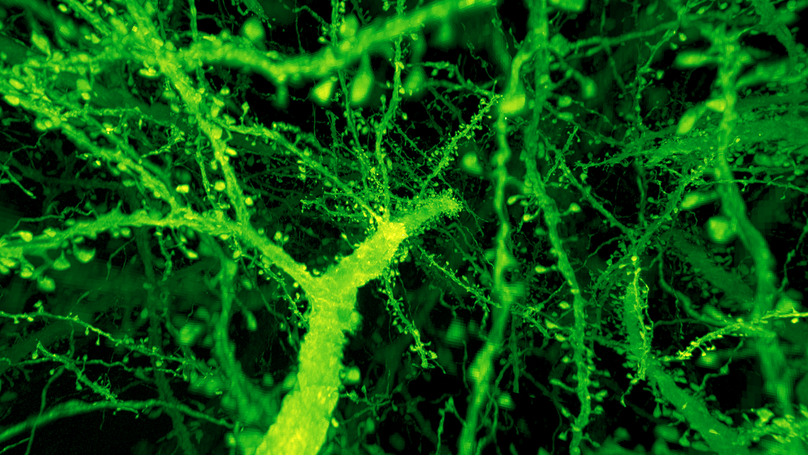
Optical and electron microscopy have made tremendous inroads toward understanding the complexity of the brain. However, optical microscopy offers insufficient resolution to reveal subcellular details, and electron microscopy lacks the throughput and molecular contrast to visualize specific molecular constituents over millimeter-scale or larger dimensions. We combined expansion microscopy and lattice light-sheet microscopy to image the nanoscale spatial relationships between proteins across the thickness of the mouse cortex or the entire Drosophila brain. These included synaptic proteins at dendritic spines, myelination along axons, and presynaptic densities at dopaminergic neurons in every fly brain region. The technology should enable statistically rich, large-scale studies of neural development, sexual dimorphism, degree of stereotypy, and structural correlations to behavior or neural activity, all with molecular contrast.

A full adult fly brain (FAFB), imaged at synaptic resolution using TEM.
- 7,062 brain slices
- 23 million images
- 115 TB on disk
- Over 1.5 years
- 1/5 acquired autonomously with custom developed Fast Stage Autoloader robot.
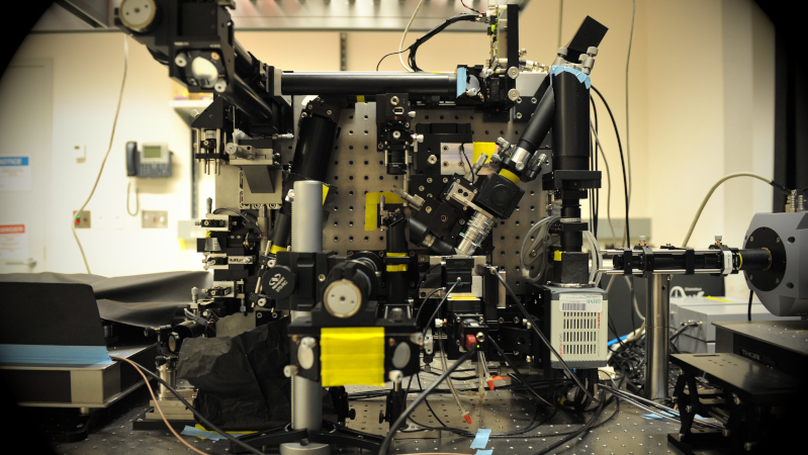

Although fluorescence microscopy provides a crucial window into the physiology of living specimens, many biological processes are too fragile, are too small, or occur too rapidly to see clearly with existing tools. We crafted ultrathin light sheets from two-dimensional optical lattices that allowed us to image three-dimensional (3D) dynamics for hundreds of volumes, often at subsecond intervals, at the diffraction limit and beyond. We applied this to systems spanning four orders of magnitude in space and time, including the diffusion of single transcription factor molecules in stem cell spheroids, the dynamic instability of mitotic microtubules, the immunological synapse, neutrophil motility in a 3D matrix, and embryogenesis in Caenorhabditis elegans and Drosophila melanogaster. The results provide a visceral reminder of the beauty and the complexity of living systems.